How do I link a table of observations and images?#
This guide shows you how to link a table of observations of any quantity with an image of the region where those observations were made. This is a specific case of linking datasets, which is explained in general in the How do I link my data in glue genes? guide. By defining these links you will be able to over-plot information about observations on the image and have selections on the image define subsets on the table and vice-versa.
Confirm your data has coordinates#
In order to link your table of observations to an image, the table must contain two columns describing the location of the observations in an image, perhaps with a scaling factor or shift/offset.
If your table also contain a set of polygons describing the region of the image covered by the observation, glue will display these regions as polygons on the image. Otherwise, glue will plot the observations as points.
Define transforms between coordinates (if necessary)#
If the pixel coordinates in your table are the same as the pixel coordinates in your image, you can follow the steps in How do I link my data in glue genes? under the If your datasets have the same attributes… to link the two datasets.
If the pixel coordinates in your table are not the same as the pixel coordinates in the image you wish to link to, you will need to define a transform between the two coordinate systems, which requires a bit of python code.
The core glue documentation explains how to set up Custom Link Functions but glue genes also provide a utility function to easily set our coordinate transforms that can be expressed as just shifts and scalings. To define these transformations you should:
1. Create (or edit) a config.py file in your current working directory. (See Configuring glue via a startup file for additional options.)
Define a tranforms by adding something like the following lines to the config.py file:
from glue.config import link_helper
from glue_genes.glue_genomics_data.image_link_functions import BaseLinearScaleLink
@link_helper(category='Images')
class Fullres_to_Hires(BaseLinearScaleLink):
description = 'Link Fullres data to HiRes data'
labels1 = ['FullRes']
labels2 = ['HiRes']
display = "Fullres <-> HiRes"
scale = 0.053972367
shift = 0
This example convert pixel coordinates in the Fullres image to pixel coordinates in the HiRes image, using a scaling factor derived from the metadata in the Space Ranger output folder. This utility function sets up a bi-directional link between the two datasets.
Use the Link Editor to define links#
Do the following steps:
1. Identify the names of the columns/components/attributes that contain the pixel locations. If you don’t know this already, create a Table Viewer to look at your data inside glue.
2. Use either the Link Data icon in the glue toolbar or the Link Data menu option in the Data Manager menu to open up the Link Editor – the interface in which you can define links between datasets.
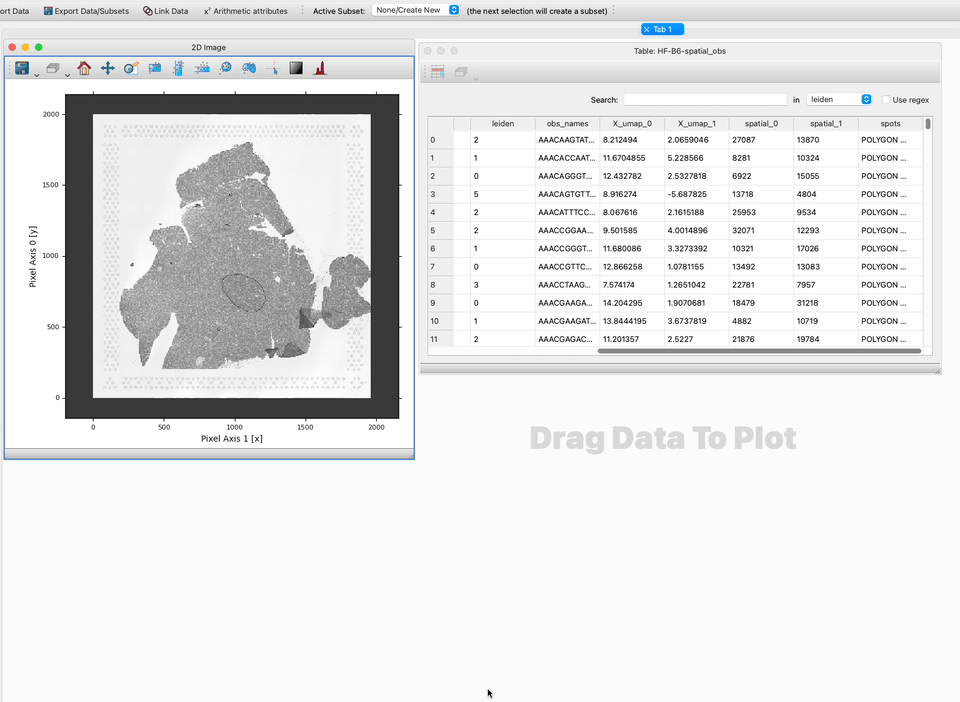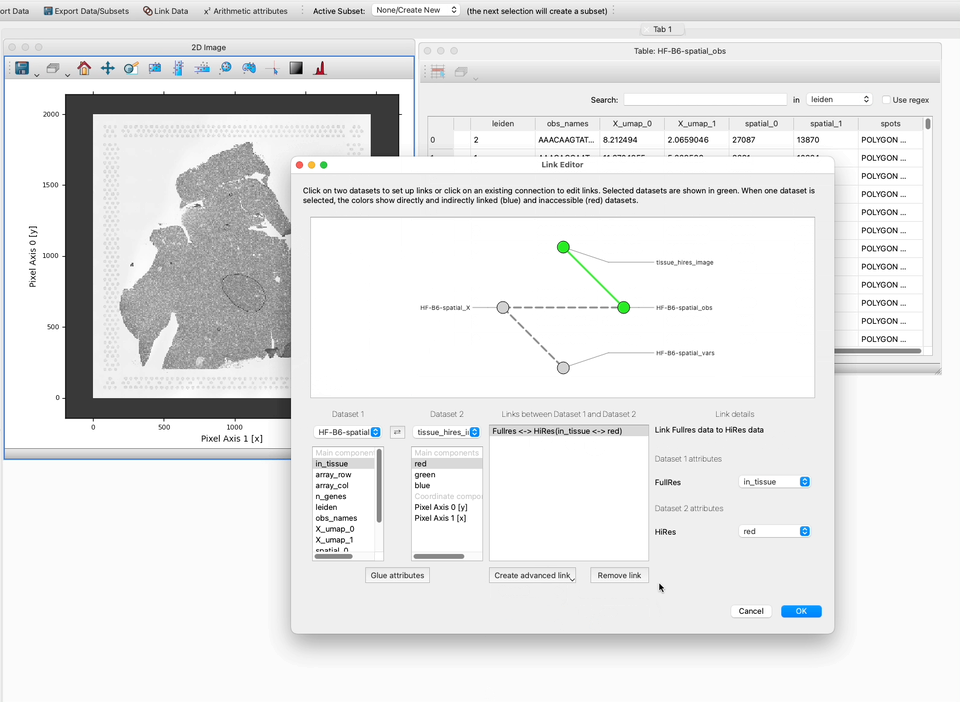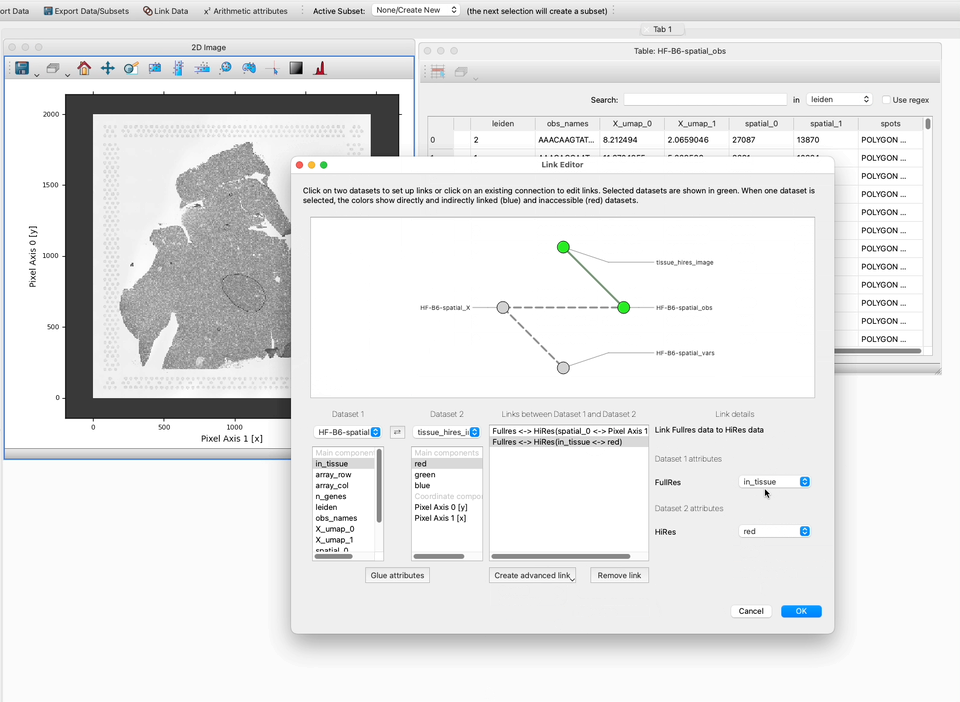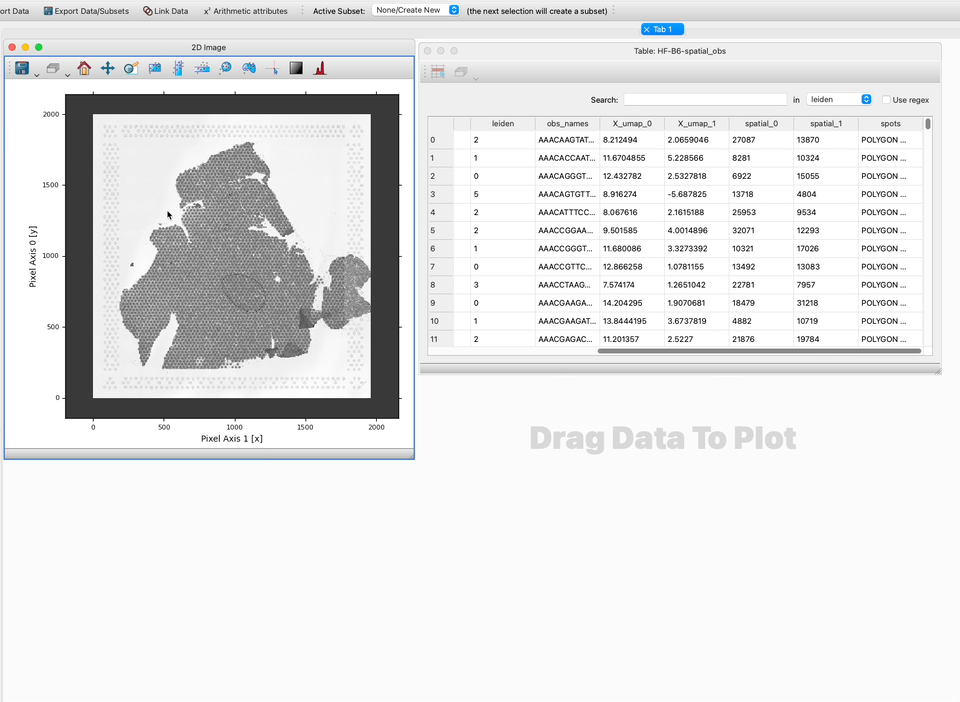
3. Select the two datasets you wish to join either by clicking on them in the central widget shows all the data sets as circles, or by selecting them in the drop downs at the left side of the Link Editor (choosing Dataset 1 and Dataset 2). Note that two datasets will be selected by default, so if you only have two datasets they will already be selected.
4. Choose the components to link on. In this case, we choose Pixel Axis 1 [x] and spatial_0 which encode the y pixel locations in the table data. When the correct components are selected, select the custom function you defined in the preceeding section of this how-to guide.
The details of the link will now show on the right-hand panel, and a solid line will show the link between the two datasets in the main display showing links between all datasets.
4. We have now joined the Y pixel coordinates. To complete this task we also need we would also need to join the Pixel Axis 0 [y] and spatial_1 attributes using our custom link function so that the table of information can be shown on top of the image.
More information#
See the section on How Data Linking Works in the glue documentation and the linking framework in the glue developer guide for more information about how glue handles links. The more general case of linking arbitrary datasets is also covered in the How do I link my data in glue genes? guide.
What next?#
Now that you have linked your data you probably want to visualize it.